Intellectual Property Portfolio
We believe innovations are the key to scientific, economic and social progress.
At the beginning of each innovation is the invention, which was fueled by human curiosity and creativity, when faced with a technical problem. In order to develop an invention into value-added applications and eventually a successful innovation is a long and difficult process, which requires strong teamwork and many resources.
baseclick has pioneered in the development of high-end nucleic acid modification for diverse applications, e.g. cell proliferation analysis and single-molecule NGS sequencing. But our relentless pursuit of perfection identifies ever new potentials for improved (click) applications, e.g. chemical engineering of mRNA. Over the past years we have created several inventions and have built an attractive patent portfolio.
As our long-term vision is to unleash the full potential of click labeling, we offer know-how and attractive licensing conditions to our partners and we would be glad to discuss licensing options with you.
Want to discuss licensing options?
Then leave a message to Dr. Thomas Frischmuth

Baseclick’s Proprietary IP Portfolio on Click Chemistry
-
New nucleic acid labelling system for the sensitive detection of analytes (WO 2006/117161 – several divisional applications)
This invention opens up new possibilities (methods, kits and reagents) for the simple and sensitive detection of samples, e.g. nucleic acids in complex biological samples using click chemistry.
-
Click chemistry for the production of reporter molecules (WO 2008/052775)
In order to increase the scope of the inventions from the previous patents, we have elaborated herein methods for the production of reporter molecules that can be used for analyte detection (e.g. nucleic acids in FISH experiments). A labeling method to introduce three different labels to the same oligonucleotides is disclosed.
-
Click chemistry on heterogeneous catalysts (WO 2010/115957)
Click reactions employing CuBr as a catalyst suffer often from instability of this Cu(I) source, which renders fresh CuBr aliquots for each reaction setup necessary. We have a simple solution: a long-term air-stable heterogeneous Cu(I) source for easy click reactions and a device to perform the reactions in.
-
Anandamide-modified nucleic acid molecules (WO 2014/009429)
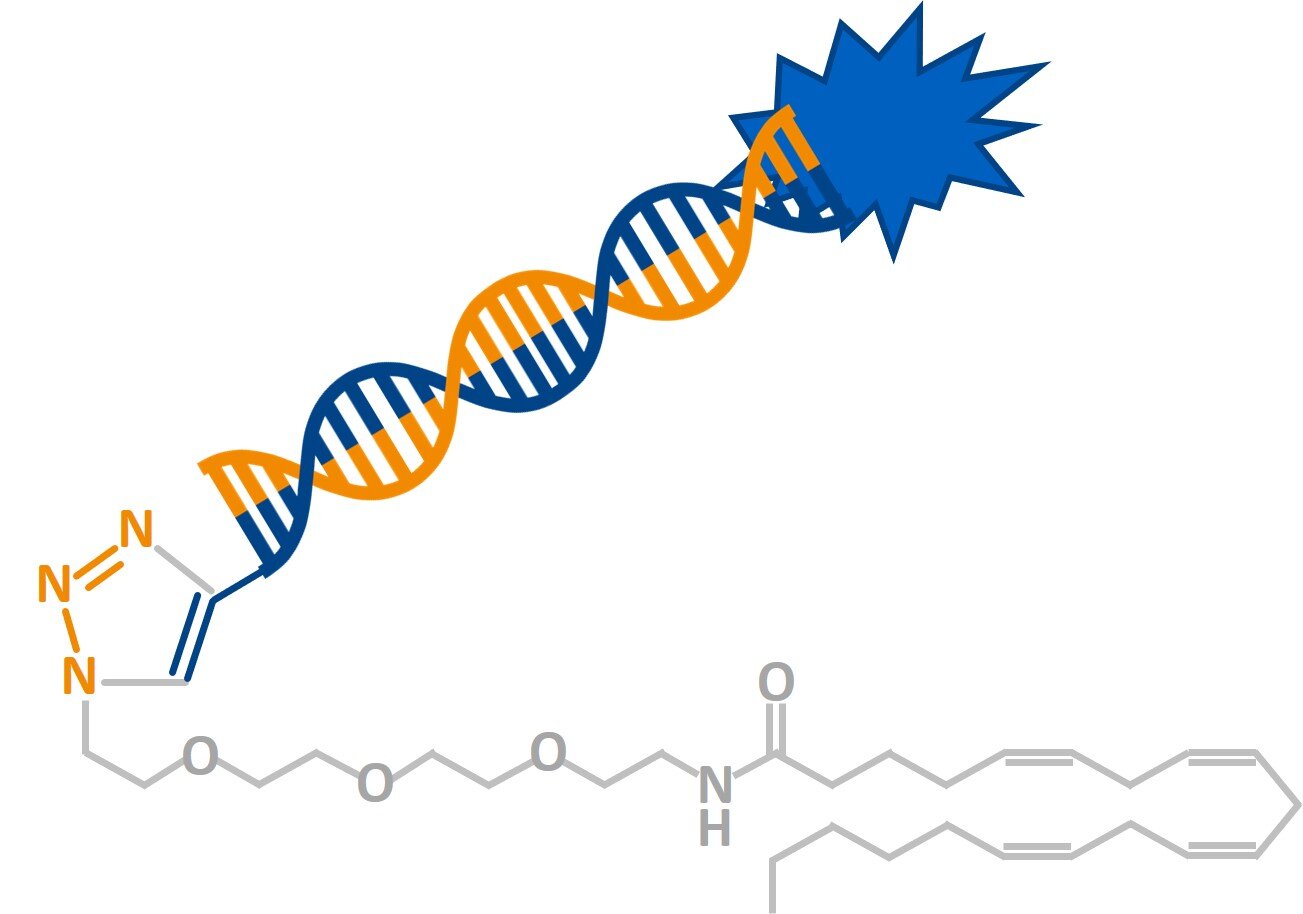
Targeted delivery of therapeutic oligonucleotides is a key challenge for this new class of drugs. Here we disclose that anandamide-modified oligonucleotides are efficiently produced via click chemistry and can be taken up by neuronal and mammalian cells for siRNA mediated gene-knockdown.
-
Saccharide-modified nucleic acid molecules (WO 2015/107115)
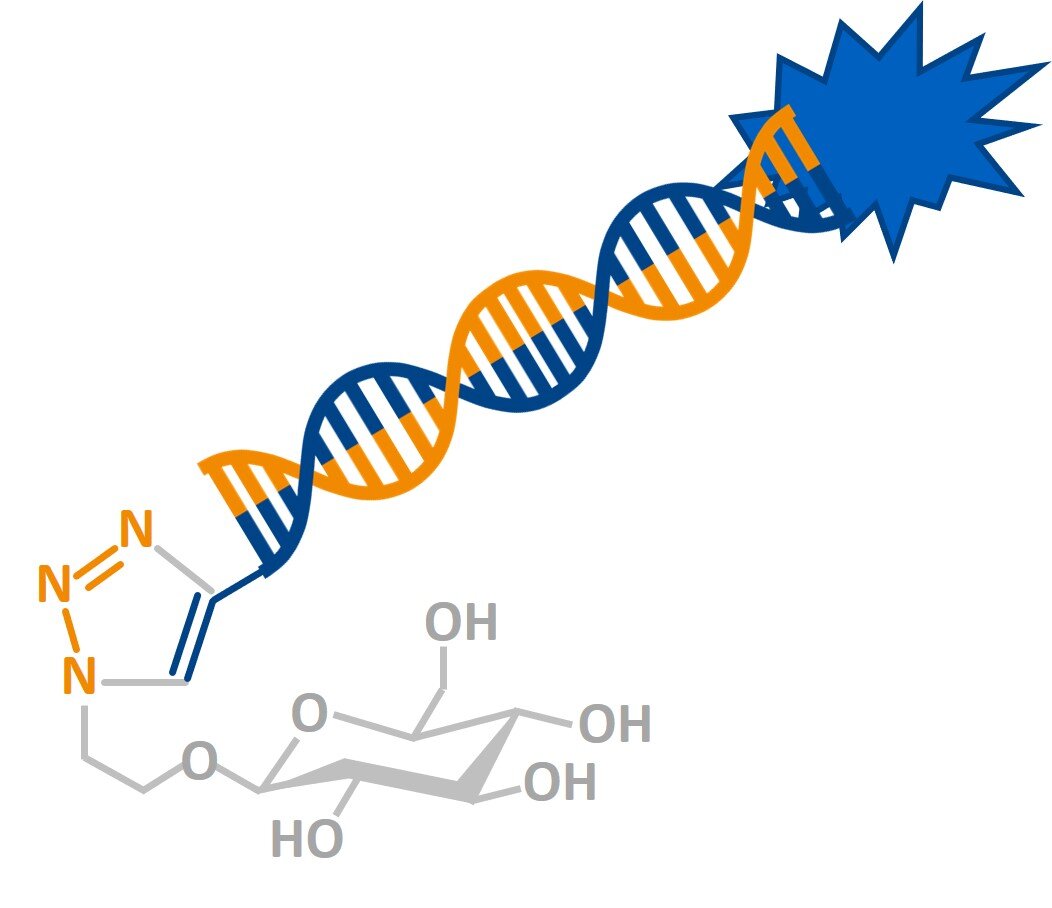
(Targeted) delivery of oligonucleotides for transient gene therapies is a major challenge. Here transfection agent free delivery of saccharide-modified oligonucleotides into plant (and mammalian cells) is disclosed. The oligonucleotide sugar conjugates are produced by click chemistry and could help to screen libraries of conjugates for a medicinal chemistry approach to RNA therapeutics.
-
Click-based ligation (WO 2019/063803)
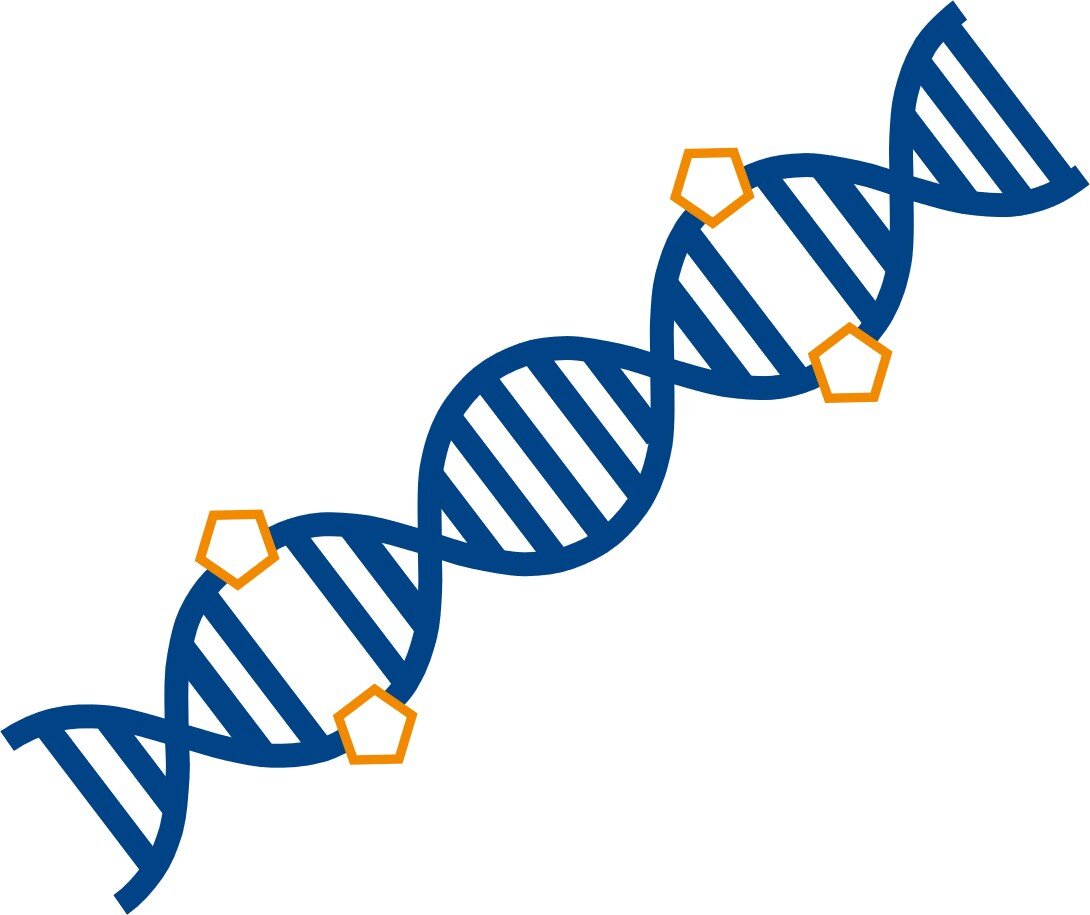
Enzymatic ligation is a key process in sample preparation for cloning and next-generation sequencing. The current ligases cannot efficiently ligate two single-stranded DNA oligonucleotides, thus limiting ligation of nucleic acid molecules mostly to double-stranded DNA. This invention discloses convenient procedures to integrate a chemical ligation protocol into current NGS library preparation workflows.
-
Click-modified mRNA (WO 2019/121803 A1)
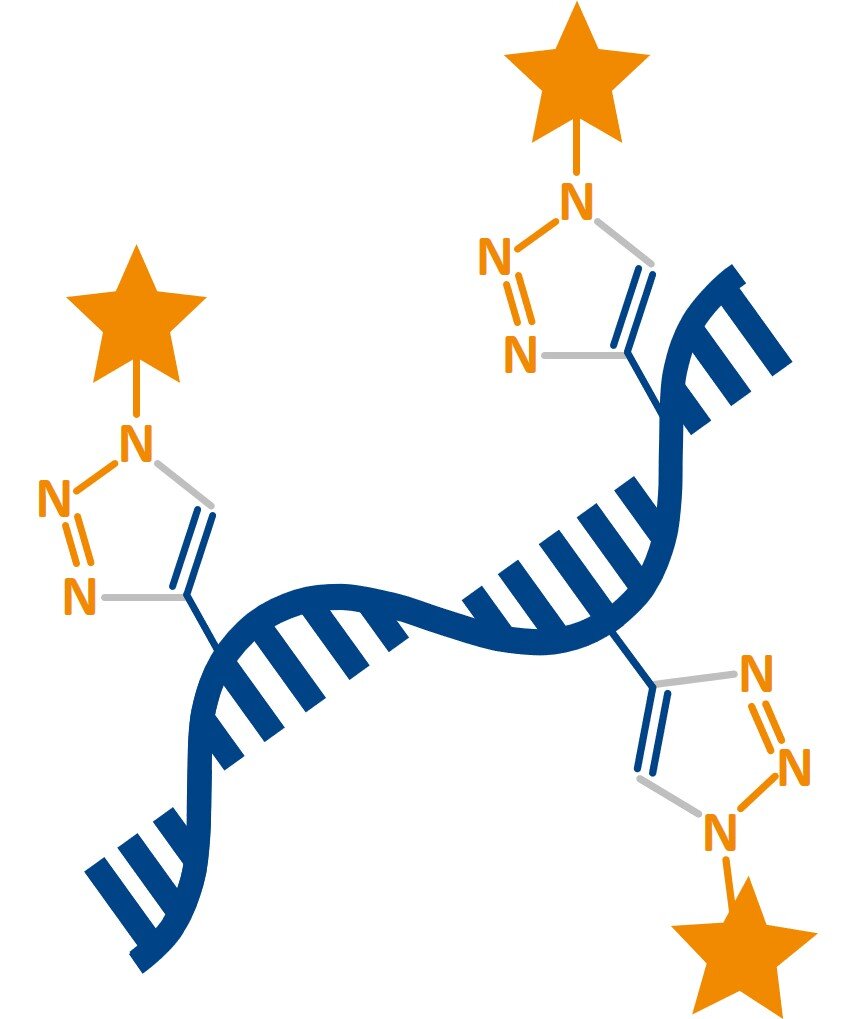
mRNAs are an emerging class of drugs with great promise, which face several challenges for commercialization. This is mostly due to a very limited capability for chemical modification during in vitro transcription of mRNA. Therefore, we developed a modular mRNA labeling method that can generate (multi-) modified functional mRNA molecules with improved properties.
-
Method of amplifying mRNAs and for preparing full length mRNA libraries
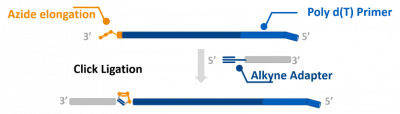
Whole transcriptome or exome sequencing has contributed enormously to understanding common and rare diseases, but has remained a challenge nonetheless. For this reason, we have developed an inventive method specifically for making 1: 1 copies of a full length RNA library for sequencing and quantitative analysis. After reverse transcription of RNA, we use click chemistry to chemically ligate adapter primers to the cDNA sequences in high yield. This allows us to amplify unknown sequences from the 3 ‘end to the 5’ end without losing information.
- Modified mRNAs for vaccine development
Trademarks:
baseclick (# 307 83 243.0/07); Eterneon (# 30 2017 013 698), Adapt Click (# 30 2018 104 831)
Want to use our trademarks?
Please let us know and we can discuss the right option for you
Copyright statement:
Texas Red® is a trademark of Molecular Probes
Cy™, Cy3™, Cy5™, Cy5.5™ and Cy7™ are trademarks of GE Healthcare

