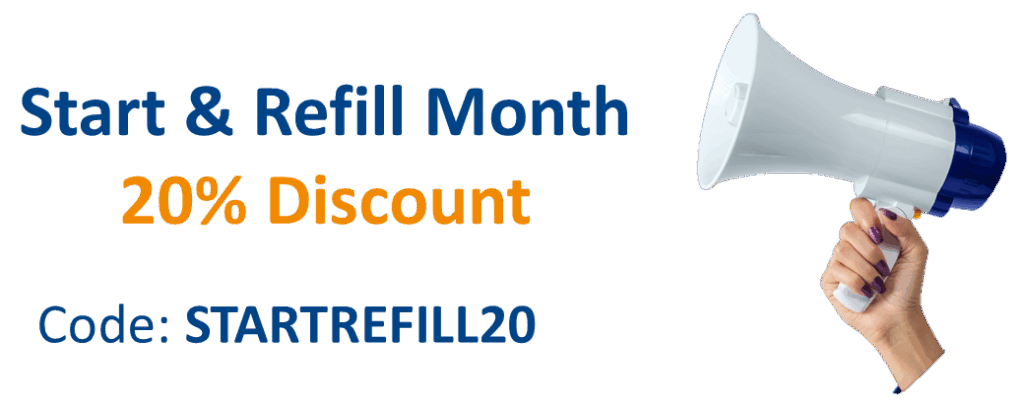

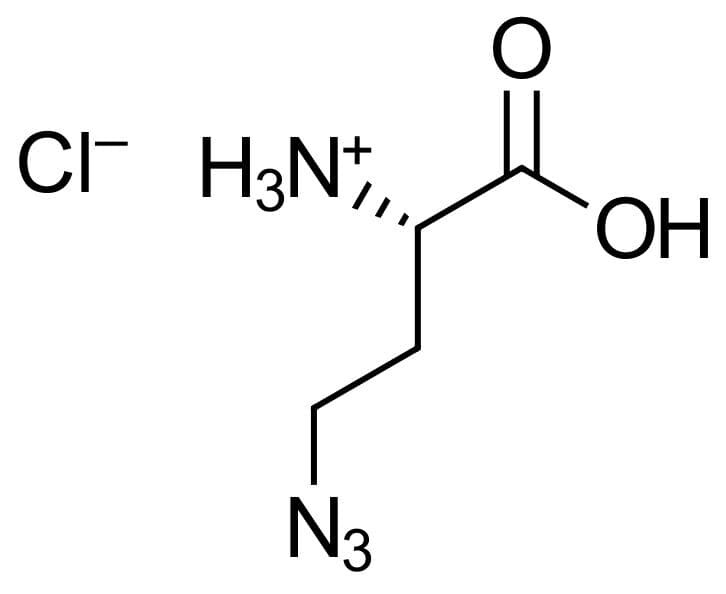
4-Azido-L-homoalanine HCl
Unnatural azido-modified amino acids for monitoring protein synthesis
| Size | Catalog No. | Price |
|---|---|---|
| 10 mg | BCAA-005-10 | € 45,00 |
| 100 mg | BCAA-005-100 | € 140,00 |
| 500 mg | BCAA-005-500 | € 400,00 |
Chemical Properties
-
Molecular Formula
C4H8N4O2 *HCI
-
Shelf Life
12 months unopened after receipt
-
Storage Conditions
2-8 °C, dry
-
Molecular Weight
144.13 g/mol * 36.45 g/mol
-
Purity
≥ 98% (HPLC)
-
Physical State
white crystalline powder
-
CAS Number
942518-29-8 (hydrochloride salt)
120042-14-0 (free acid) -
Solubility
soluble in e.g. 0.1 M NaOHaq
-
Additional name
(S)-2-Amino-4-azidobutanoic acid hydrochloride, 4-Azido-L-homoalanine hydrochloride, H-L-Aha-OH*HCl
Product Information
Bioorthogonal Protein Labeling for Advanced Molecular Engineering
4-Azido-L-homoalanine hydrochloride ((S)-2-Amino-4-azidobutanoic acid hydrochloride) is a synthetic, unnatural amino acid based on homoalanine carrying an azide group for site-specific protein modification and bioorthogonal labeling via Click Chemistry (SPAAC / CuAAC) or Staudinger ligation.
This reagent can be incorporated into peptides or proteins through:
- Genetic Code Expansion: Engineered tRNAs (based on prokaryotic tRNAs) recognize amber codons (usually UAG) and incorporate 4-Azido-L-homoalanine at a defined position during biosynthesis.
- Solid Phase Peptide Synthesis (SPPS): Enables chemical incorporation at any position in the peptide sequence for versatile functionalization.
Once incorporated, the azide moiety reacts selectively with alkyne-modified partners (DBCO, BCN, PEGs, dyes) or P(III)-containing reagents, enabling efficient conjugation without side reactions.
Why choose 4-Azido-L-homoalanine?
Traditional NHS esters react with all primary amines available (e.g., lysines), leading to:
- Non-specific labeling at multiple sites
- Requires basic pH (8–9) and overnight reactions
- Risks for protein degradation and structural changes
In contrast, azide-based labeling with 4-Azido-L-homoalanine offers:
- Site-specific modification at the planned position
- Fast reactions at neutral pH with high yields
- Preserves tertiary structure
- Fully biorthogonal, no interference with native functional groups
- Lower reagent consumption and atom-economic click chemistry for a cost-effective, eco-friendly workflow
Applications of 4-Azido-L-homoalanine HCl
4-Azido-L-homoalanine is widely used for protein and peptide labeling, enabling precise site-specific modification through bioorthogonal click chemistry. It is an excellent choice for antibody modification and ADC synthesis, supporting the development of targeted therapeutics. Researchers also use this reagent for linking targeting moieties to proteins in drug delivery and diagnostic applications. Additionally, its compatibility as a building block for solid-phase peptide synthesis (SPPS) makes it ideal for custom peptide design in pharmaceutical development, biotechnology, and advanced biochemistry workflows.
Therefore, 4-Azido-L-homoalanine provides a highly selective, biocompatible, and stable way to perform peptide/protein labeling in a bioorthogonal way. This enables precise molecular engineering for pharmaceutical development and advanced biotechnology or biochemistry research, without compromising the structure and function of the peptide/protein. The functionalization by Staudinger Ligation is also possible in a biorthogonal way.
Key Benefits
The main advantages of 4-Azido-L-homoalanine are:
- Integrity of tertiary structure: SPAAC and CuAAC reactions can be performed at neutral pH in short reaction times and, therefore, chemical modification of your peptide/protein is possible without the risk of degradation of the target biomolecule.
- Fast & Efficient Click Reactions: The used click chemistry forms highly stable triazole bonds with all alkynes or alkyne derivates (e.g. DBCO, BCN) which can be used for strain promoted alkyne-azide cycloaddition in short reaction time.
- Specific labeling: Since 4-Azido-L-homoalanine is incorporated at an exact defined position of your peptide/protein highly specific labeling at this position of your peptide/protein is possible.
LITERATURE
Expanding One-Pot Cell-Free Protein Synthesis and Immobilization for On-Demand Manufacturing of Biomaterials, A. Benitez-Mateos et al., 2018, ACS Synthetic Biology, Vol. 7(3), p. 875-884.
https://doi.org/10.1021/acssynbio.7b00383
Bio-orthogonal labeling as a tool to visualize and identify newly synthesized proteins in Caenorhabditis elegans, M. Ullrich et al., 2014, Nature Protocols, Vol. 9(9), p. 2237–2255.
https://doi.org/10.1038/nprot.2014.150
Ubiquitin C-terminal hydrolase L1 (UCH-L1) loss causes neurodegeneration by altering protein turnover in the first postnatal weeks, A. T. Reinicke et al., 2019, Proceedings of the National Academy of Sciences, Vol. 116(16), p. 7963-7972.
https://doi.org/10.1073/pnas.1812413116
Exclusive enteral nutrition initiates individual protective microbiome changes to induce remission in pediatric Crohn’s disease, Häcker, D et al., 2024, Cell host & microbe, 32(11), 2019-2034.
https://doi.org/10.1016/j.chom.2024.10.001
FAQ
-
Is it possible to generate azide- or alkyne-modified peptides?
With modified amino acids azide- or alkyne-modified peptides can be prepared by solid-phase synthesis.
-
How can de novo protein biosynthesis be monitored?
De novo protein biosynthesis can be monitored by feeding of metabolite analogues (so-called metabolic labeling) and subsequent click reaction. Azido-homoalanine for example is recognized as a methionine analogue and is incorporated into de novo synthesized proteins in methionine-free medium conditions. The resulting proteins contain azide moieties and thus can be detected after click to an alkyne-containing reporter molecule (e.g. a fluorescent dye). This non-radioactive method has major practical advantages compared to traditional 35S amino acid incorporation methods.
Alternatively, O-propargyl-puromycin is efficiently incorporated into proteins during de novo protein biosynthesis and can be used in complete medium. The resulting alkyne protein fragments can be detected via click to azide-containing reporter molecules. -
What click conditions should be used for protein click reactions?
A catalyst system based on CuSO4 and sodium ascorbate is recommended in combination with dye azides to label alkyne-modified proteins. Please also refer to our general Click protocols for more details.
Due to the 20 (21) amino acids that are the building blocks of proteins, the physicochemical properties of proteins are more diverse compared to oligonucleotides, which are just composed of 5 major building blocks. Therefore, finding the optimal click conditions is more difficult compared to oligonucleotides and labeling rates are usually lower. Please note that despite these difficulties detection applications (e.g. de novo protein biosynthesis detection) are easily feasible. -
Why use 4-Azido-L-homoalanine instead of NHS ester chemistry?
It provides precise, site-specific labeling under mild conditions, avoiding multiple modification sites and preserving protein integrity.
-
Which click reactions are supported?
SPAAC (copper-free) and CuAAC (copper-catalyzed) for rapid, efficient conjugation.
-
Can it be used in SPPS?
Yes, it can be incorporated at any position during peptide synthesis.
-
What labeling partners are compatible?
DBCO/BCN/primary alkyne-modified PEGs, fluorescent dyes, antibodies, and targeting moieties.

