
EdU T Cell Proliferation Assay for Flow Cytometry
Accurate, Direct and Multiparametric Analysis of Human T-cell Proliferation

| Size | Catalog No. | Price |
|---|---|---|
| Dye 488 / 48 Assays | BCK-TCell-FC488_48 | € 170,00 |
| Dye 488 / 192 Assays | BCK-TCell-FC488_192 | € 400,00 |
| Dye 647 / 48 Assays | BCK-TCell-FC647_48 | € 170,00 |
| Dye 647 / 192 Assays | BCK-TCell-FC647_192 | € 400,00 |
Chemical Properties
-
Shelf Life
12 months unopened after receipt
-
Storage Conditions
2-8 °C
-
Physical State
kit system made of different components
-
CAS Number
n.a.
-
Excitation (max)
Dye 488: 496 nm | Dye 647: 643 nm
-
Emission (max)
Dye 488: 516 nm | Dye 647: 662 nm
-
Ɛ (max)
Dye 488: 83.000 cm-1M-1 | Dye 647: 250.000 cm-1M-1
-
Preparation/Handling
please see user manual of the kit
Product Information
High-resolution detection of T-Cell Proliferation with ClickTech EdU Kit
The EdU T Cell Proliferation Assay for Flow Cytometry is a powerful tool for the direct detection of DNA synthesis in activated human T cells via click chemistry-based EdU incorporation. Unlike dye-dilution methods, this kit precisely detects cells actively synthesizing DNA during S-phase of the cell cycle, providing high-resolution data even in complex co-culture or immunotherapy-relevant systems.
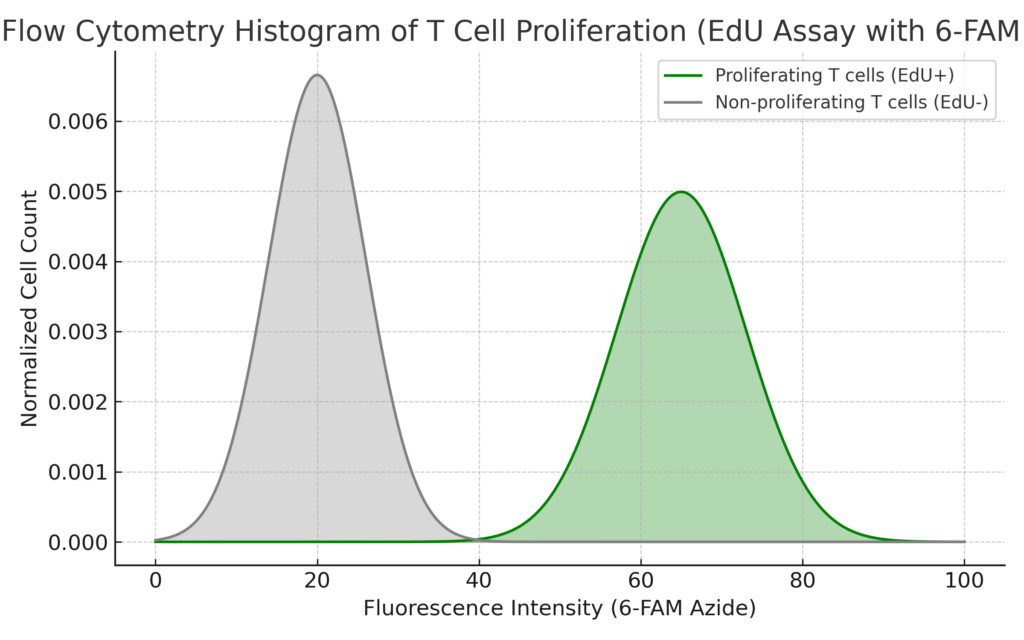
Figure 1: Flow cytometry histogram showing T cell proliferation detected via EdU incorporation and 6-FAM azide staining. The histogram displays two distinct populations:
Proliferating T cells (EdU⁺) are represented by the larger peak at higher fluorescence intensity (colored in light green), indicating successful incorporation of EdU during DNA synthesis.
Non-proliferating T cells (EdU⁻) form a separate peak at lower fluorescence intensity (gray), reflecting cells that did not enter S-phase during the EdU pulse.
Key Features
- Direct detection of proliferating T cells through EdU incorporation
- Ideal for immunotherapy, vaccine research, T-cell manufacturing, and drug screening
- No need for harsh DNA denaturation steps (unlike BrdU)
- Multiplex with surface and intracellular markers for in-depth immune profiling
Applications
- Monitoring T-cell activation and proliferation post-stimulation (e.g., CD3/CD28)
- Characterizing antigen-specific T-cell responses
- Assessing proliferation in chimeric antigen receptor T-cell therapy (CAR-T) and checkpoint inhibitor studies
- Evaluating donor–recipient responses in Mixed Lymphocyte Reactions (MLR)
- Tracking T-cell expansion in bioreactors and ex vivo cultures
- High-content immunophenotyping with transcription factors or cytokines
Why EdU instead of dye dilution?
Dye dilution methods like CellTrace™ Far Red or CFSE are commonly used to track T-cell divisions. However, they can suffer from:
• Inconsistent staining in complex populations
• Signal loss over time, complicating long-term studies
• Interference with cell function or downstream staining
The EdU T Cell Proliferation Assay for Flow Cytometry offers a reliable, DNA-level readout of cell proliferation unaffected by cell type, metabolic state, or dye efflux.
Compare with other proliferation assays
| Method | Sensitivity | Multiplexing | Harsh Processing | Temporal Resolution |
| EdU | ● ● ● ● ○ | ● ● ● ● ● | No | High (S-phase specific) |
| BrdU | ● ● ● ● ○ | ● ● ○ ○ ○ | Yes (DNA denaturation) | Moderate |
| Ki-67 | ● ● ● ○ ○ | ● ● ● ● ○ | No | Broad (any active phase) |
| CellTraceTM | ● ● ● ● ○ | ● ● ● ○ ○ | No | Division-based, cumulative |
Kit components
- EdU reagent (optimized for T-cell incorporation)
- ClickTech detection buffer system
- Permeabilization & fixation buffers
- Detailed protocol optimized for human T-cell activation workflows
Workflow overview
- Stimulate T cells (e.g., anti-CD3/CD28, antigens, APCs)
- Add EdU to culture medium
- Incubate (typically 4–24 hours)
- Harvest and stain cells using our optimized click chemistry protocol
- Analyze via flow cytometry (can be multiplexed with CD3, CD4, CD8, CD25, FoxP3, IFN-γ, etc.)
Available Formats
- 48 tests
- 192 tests
- Bulk formats available for CROs and manufacturing partners
Ready to upgrade your T-cell assays?
The ClickTech EdU T-cell Proliferation Kit offers clear, direct insight into T-cell dynamics without the limitations of dilution dyes. Perfect assay for researchers in cancer immunology, infectious disease, autoimmunity, or cell therapy development.
LITERATURE
A sensitive and less cytotoxic assay for identification of proliferating T cells based on bioorthogonally-functionalized uridine analogue, F. C. Stempels et al., 2022, Journal of Immunological Methods, Vol. 502, p. 113228.
https://doi.org/10.1016/j.jim.2022.113228
FAQ
-
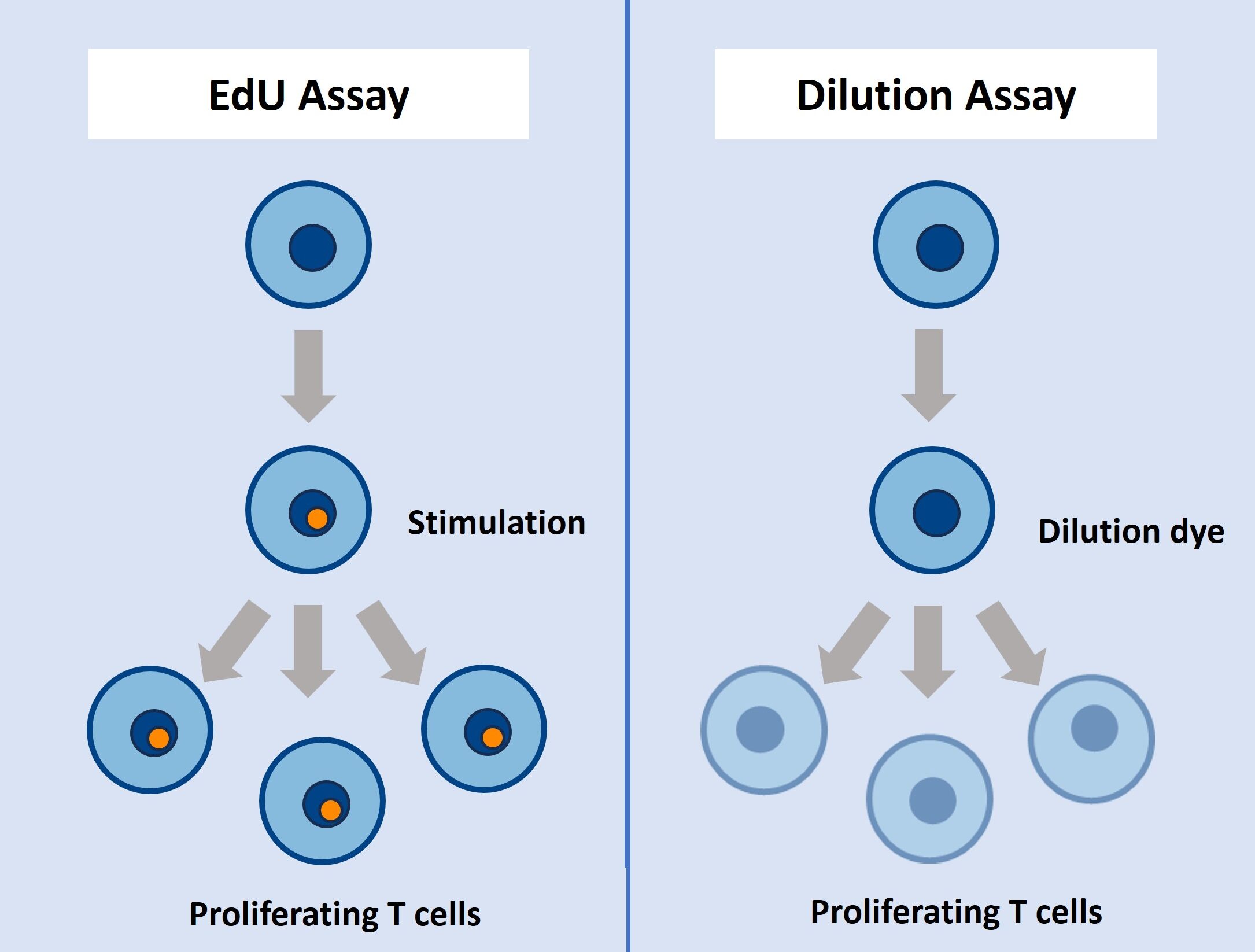
How does EdU labeling compare to methods based on label dilution?
Mostly, dilution-based T cell proliferation probes conjugate unspecifically to amine groups in cells. Each cell division then results in dilution (takes several days) of the probes. Main disadvantages are cytotoxicity, an impact on T cell activation and long incubation time (several days and up to a week) before readout. The superior EdU cell proliferation assay relies on the incorporation of the modified uridine analog EdU in the DNA of proliferating T cells (just 2 hours before harvesting) and subsequent detection. Both methods enable detection and quantification of T cell proliferation by flow cytometry. Our EdU incorporation assay is very sensitive, and can be multiplexed with other fluorescent stainings.
- EdU outperforms a standard dilution-based probe in detecting T cell proliferation
- The EdU assay is less cytotoxic for human T cells
- The EdU assay offers superior signal-to-background ratio
- The EdU assay allows for better discernable interferon gamma responses
-
Can I combine the EdU T cell Proliferation assay and staining with PerCP, APC, APC-based tandems, RPE, PR-tandem, Quantum Dot antibody conjugates or intracellular antigens?
Yes, this is feasible. Please note that staining with PerCP, APC and APC-based tandems may be performed before the EdU detection step (click reaction), while RPE, PR-tandem or Quantum Dot antibody conjugates and detection of intracellular antigens should be performed after the click reaction. Check also the user manual for further information. Please, make sure that the emission spectra of the fluorochromes to be combined do not overlap extensively, so that their signals can be distinguished unequivocally using appropriate emission filters or spectral demixing.
-
When may break points be introduced during the protocol?
It is possible to safely interrupt the protocol after the fixation step. Thereto, remove the fixation solution and wash as suggested by the user manual, then the cells can be stored in buffer at 4° C. Alternatively, the experiment can also be safely interrupted after permeabilization, again as described above.
Please note: It is important to proceed with the experiment if the click cocktail for the detection of the EdU has been prepared already. -
How to determine the EdU incubation time?
The EdU incubation time depends on the cell type or organism, the applied EdU concentration and the experimental design. For a start it is advisable to refer to a literature protocol (which is close to your experimental setup) and to test the conditions with a low number of samples. As a general guideline we recommend to use a maximum of 10 µM final EdU in the cell culture medium for incubations. For longer incubation (> 1 day) the concentration should be decreased to 1-5 µM.

