
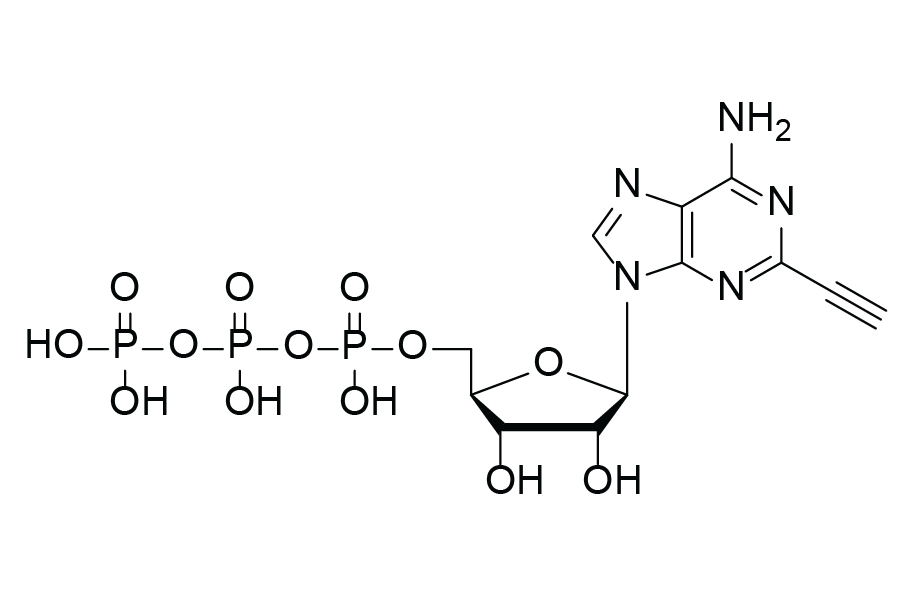
2-Ethynyl-ATP (2-EATP)
Triphosphate for modifying of 3'END RNA
| Size | Catalog No. | Price |
|---|---|---|
| 1 µmol | BCT-30-S | € 190,00 |
| 5 µmol | BCT-30-L | € 720,00 |
Chemical Properties
-
Molecular Formula
C12H16N5O13P3
-
Shelf Life
12 months unopened after receipt
-
Storage Conditions
-20 °C, dark, dry
-
Molecular Weight
531.20 g/mol
-
Purity
≥ 90% (HPLC)
-
Physical State
colorless to slightly yellow solution (10 mM)
-
CAS Number
1802333-53-4
-
Absorption (max)
λmax = 265 nm
-
Ɛ (max)
10.600 cm-1M-1
Product Information
Alkyne‑Modified ATP for Click‑Enabled RNA Labeling & ATP‑Dependent Enzyme Studies
2‑Ethynyl‑ATP (2‑EATP) is a terminal‑alkyne–modified adenosine‑5′-triphosphate designed for biochemical assays, RNA labeling workflows, and bioorthogonal click chemistry. Its reactive ethynyl group enables highly efficient copper(I)‑catalyzed azide‑alkyne cycloaddition (CuAAC), allowing fluorescent dyes, biotin labels, affinity handles, or biomolecules to be covalently attached.
2‑EATP is widely used in ATP‑binding studies, ATP‑dependent enzyme assays, in vitro RNA polyadenylation, and advanced RNA imaging and quantification techniques.
Key Features
- Alkyne-functionalized ATP analog for CuAAC click chemistry
- Suitable for ATP-binding assays, kinase studies, ATPase assays, and enzyme kinetics
- Efficiently incorporated by recombinant poly(A) polymerase for RNA 3’-end labeling
- Enables flexible downstream modification with:
- Biotin‑azide
- Fluorophore‑azides (488, 555, 594, 647)
- Azide‑linked crosslinkers
- High purity (≥ 90%, HPLC)
- Supplied as a ready‑to‑use 10 mM solution
- Long shelf life (12 months at –20 °C, dry, dark)
How 2‑Ethynyl‑ATP Works
- Enzymatic Incorporation: 2‑EATP is recognized by several ATP‑dependent enzymes including:
-
- Recombinant poly(A) polymerase
- Kinases
- ATPases
- RNA‑processing enzymes
This enables biochemical characterization of ATP‑dependent mechanisms.
- RNA Polyadenylation and 3’-End Labeling
During in vitro polyadenylation, 2‑EATP is incorporated into nascent poly(A) tails, generating click‑reactive RNA molecules suitable for downstream functionalization.
- CuAAC Click Chemistry Functionalization
The terminal alkyne reacts with azides to attach:
-
- Biotin (for purification workflows)
- Fluorophores (for RNA imaging)
- Protein conjugates (for crosslinking analyses)
Applications
- ATP‑Dependent Enzyme Studies
Ideal for investigating:
-
- ATP‑binding proteins
- Kinases
- ATPases
- Enzymatic mechanisms relevant to cancer and neurological disorders
- In Vitro Polyadenylation of RNA
Enables:
-
- Functionalized RNA 3’-end labeling
- Poly(A)-profiling
- RNA Tail engineering for mechanistic studies
- Click‑Enabled RNA Labeling and Purification
Allows downstream introduction of:
-
- Biotin tags
- Fluorescent dyes
- Bioorthogonal linkers
- for imaging, pull‑down assays, or conjugations.
- Advanced RNA Imaging & Analysis Technologies
Used in high‑impact methods such as:
-
- Rolling FISH (rFISH)
- Single‑cell polyadenylation profiling
- Poly(A)-trap assays
LITERATURE
Click-encoded rolling FISH for visualizing single-cell RNA polyadenylation and structures, F. Chen et al., 2019, Nucleic Acids Research, Volume 47, Issue 22, Page e145.
https://doi.org/10.1093/nar/gkz852
Global profiling of stimulus-induced polyadenylation in cells using a poly(A) trap, D. Curanovic et al., 2013, Nat Chem Biol, Vol. 9, p. 671–673.
https://doi.org/10.1038/nchembio.1334
Template-independent enzymatic synthesis of RNA oligonucleotides, D. J. Wiegand et al., 2025, Nature Biotechnology, Vol. 43(5), p. 762–772.
https://doi.org/10.1038/s41587-024-02244-w
Click-encoded rolling FISH for visualizing single-cell RNA polyadenylation and structures, F. Chen et al., 2019, Nucleic Acids Research, Vol. 47(22), p. e145.
https://doi.org/10.1093/nar/gkz852
Global profiling of stimulus-induced polyadenylation in cells using a poly(A) trap, D. Curanovic et al., 2013, Nature Chemical Biology, Vol. 9, p. 671–673.
https://doi.org/10.1038/nchembio.1334
A Platform for Controlled Template-Independent Enzymatic Synthesis of RNA Oligonucleotides and Therapeutics, D. J. Wiegand et al., 2023, bioRxiv, 2023-06.
https://doi.org/10.1101/2023.06.29.547106
FAQ
-
Which enzymes accept 2‑EATP as a substrate?
It is processed by poly(A) polymerase, kinases, ATPases, and various ATP‑dependent enzymes.
-
Can 2‑EATP be used in click chemistry?
Yes, the alkyne enables efficient CuAAC reactions with azide‑functionalized dyes, biotin, or proteins.
-
How should the product be stored?
Store at –20 °C, protected from light. Short-term exposure to room temperature (up to one week) is tolerated.
-
Is the product RUO?
Yes, 2‑EATP is designated Research Use Only (RUO).

