
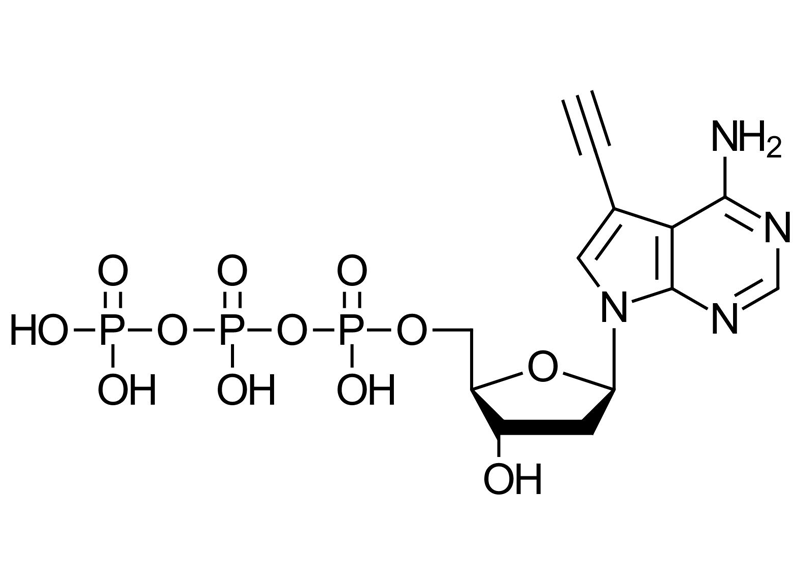
Alkyne-dATP (7-deaza-7-Ethynyl-dATP)
Modified triphosphate for incorporation in PCR reaction
| Size | Catalog No. | Price |
|---|---|---|
| 1 µmol | BCT-21-S | € 120,00 |
| 5 µmol | BCT-21-L | € 380,00 |
Chemical Properties
-
Molecular Formula
C13H17N4O12P3
-
Shelf Life
12 months unopened after receipt
-
Storage Conditions
-20 °C
-
Molecular Weight
514.22 g/mol
-
Purity
≥ 98% (HPLC)
-
Physical State
10 mM clear colorless to yellow solution
-
CAS Number
1208963-36-3 (free acid)
-
Absorption (max)
λmax = 278 nm
-
Additional name
Alkyne-deoxyadenine-5’-triphosphate
Product Information
Modified Triphosphate for PCR Incorporation & Click‑Enabled DNA Functionalization
Alkyne‑dATP (7‑deaza‑7‑Ethynyl‑dATP) is a chemically modified deoxyadenosine‑5’-triphosphate containing a terminal alkyne at the C7 position of the purine ring. This modification enables efficient incorporation into DNA during PCR and allows robust downstream functionalization via copper(I)‑catalyzed azide‑alkyne cycloaddition (CuAAC).
The nucleotide is accepted by high‑fidelity polymerases such as Pwo, Deep Vent exo‑, and KOD XL, allowing simple substitution of 1–10% of natural dATP. It also improves sequencing and PCR performance on GC‑rich or highly structured templates by reducing secondary structure formation.
The 7‑deaza modification replaces the nitrogen at the N7 position of the purine ring with a carbon atom, which suppresses Hoogsteen base pairing and prevents polymerase stalling during PCR or sequencing.
The resulting DNA fragments carry a stable, bioorthogonal alkyne handle suitable for fluorescent labeling, biotinylation, surface conjugation, and other advanced molecular biology applications.
Key Features
- Alkyne‑modified dATP enabling Cu(I)‑catalyzed azide‑alkyne cycloaddition (CuAAC)
- High incorporation efficiency using polymerases such as Pwo, Deep Vent exo‑, and KOD XL
- Produces stable, clickable PCR amplicons ready for downstream labeling
- Recommended substitution: 1–10% of dATP in PCR
- High purity: ≥ 98% (HPLC)
- Supplied as 10 mM colorless-to-yellow solution
- Long shelf life: 12 months at –20 °C
- deal for DNA labeling, molecular biology assays, and nucleic acid engineering
How Alkyne‑dATP Works
- PCR Incorporation
During PCR, Alkyne‑dATP substitutes for natural dATP with high efficiency. Polymerases incorporate the nucleotide into the DNA backbone, placing the alkyne handle at the exact positions of adenine sites.
- Bioorthogonal Click Functionalization
The incorporated alkyne reacts selectively with azide-functionalized labels using CuAAC click chemistry, enabling attachment of:
-
- Fluorophores
- Biotin and affinity tags
- Small molecules and chemical handles
- Benefits of the 7‑Deaza Modification
The 7‑deaza substitution:
-
- Alters the electronic environment of the purine ring
- Reduces unwanted Hoogsteen pairing
- Prevents formation of secondary structures
- Improves polymerase recognition and bypass of structured DNA regions
Applications of Alkyne‑dATP
- Click‑Enabled DNA Labeling
-
- Fluorescent labeling
- Biotinylation for pull‑downs
- Conjugation to peptides, proteins, or nanoparticles
- PCR Probe Synthesis
Allows generation of clickable DNA fragments for use in:
-
- Molecular diagnostics
- FISH probe backbone synthesis
- Hybridization assays
- Structural DNA Studies
The 7‑deaza core reduces formation of secondary structures such as:
-
- G‑quadruplexes
- Hoogsteen base pairs
- Improving reproducibility in difficult PCR regions.
- Next‑Generation Sequencing & DNA Engineering
-
- Improved sequencing through structured templates
- Template engineering for click‑based barcoding
LITERATURE
5-Substituted Pyrimidine and 7-Substituted 7-Deazapurine dNTPs as Substrates for DNA Polymerases in Competitive Primer Extension in the Presence of Natural dNTPs, H. Cahova et al., 2016, ACS Chem. Biol., Vol. 11(11), p. 3165–3171.
https://doi.org/10.1021/acschembio.6b00714
Direct voltammetric analysis of DNA modified with enzymatically incorporated 7‑deazapurines, H. Pivonková et al., 2010, Analytical Chemistry, Vol. 82(16).
https://doi.org/10.1021/ac100757v
FAQ
-
How much natural dATP should be replaced with Alkyne‑dATP?
We recommend substituting 1–10% of dATP in PCR reactions.
-
Which polymerases efficiently incorporate Alkyne‑dATP?
Pwo, Deep Vent exo‑ and KOD XL polymerases incorporate it well.
-
Can the modified DNA be used for click chemistry?
Yes. The incorporated alkyne enables CuAAC attachment of dyes, biotin, or other azide‑labeled molecules.
-
Is this product RUO?
Yes, it is for Research Use Only (RUO).

