
DNA-ClickSeq Library Prep Kit with dual indexing
Fragmentation‑free, ligation‑free Illumina® library prep with robust dual indexing

| Size | Catalog No. | Price |
|---|---|---|
| 12 rxn | BCK-DNAseqDual | € 396,00 |
Chemical Properties
-
Shelf Life
12 months unopened after receipt
-
Storage Conditions
– 20 °C
-
Physical State
kit system made of different components
-
CAS Number
n.a.
-
Preparation/Handling
please see user manual of the kit
Product Information
Technical Overview
DNA‑ClickSeq with Dual Indexing streamlines Illumina‑compatible library construction by replacing enzymatic adapter ligation and mechanical fragmentation with click chemistry (CuAAC). Stochastic 3′‑azido termination creates clean, azido‑terminated cDNA fragments that are chemically ligated to alkyne‑adaptors, then PCR‑completed with unique i5/i7 indices primer for accurate multiplexing. The result is a faster, simpler, and lower‑bias workflow, ideal for high‑throughput genomics and applications where chimera suppression matters [1] [2].
Why DNA‑ClickSeq
- Fragmentation‑free, ligation‑free: Eliminates two major sources of bias and artifacts, accelerating time‑to‑sequence [1] [2].
- Ultra‑low chimera rate: Literature reports fewer than 3 aberrant recombination events per million reads, enabling confident detection of rare events and structural changes [1].
- Robust dual indexing (i5/i7): Minimizes index hopping and mis‑assignment; scales multiplexing without compromising data integrity.
- Uniform coverage, low error: Produces unbiased libraries with error rates comparable to standard methods while avoiding fragmentation‑induced bias [1].
- Polymerase‑compatible triazole linkage: The CuAAC‑formed triazole is biocompatible with RT/DNA polymerases and supports efficient PCR amplification [2].
- Fast & flexible: Complete the workflow in ~6 hours with 10 ng–4 µg DNA input (≥100 ng recommended for best complexity).
How it Works (Technical Overview)
- Azido‑Termination: Reverse transcription on DNA templates with a 6N primer (partial p7) in the presence of 3′‑azido‑2′,3′‑dideoxynucleotides (AzNTPs) stochastically terminates extension, yielding 3′‑azido‑terminated cDNA fragments [1].
- Click‑Ligation (CuAAC): 3′‑azido fragments are click‑ligated to p5‑adaptor oligos carrying a 5′ alkyne, creating a stable triazole linkage without ligases [1] [2].
- Dual‑Index PCR: PCR completes p5/p7 sequences and adds distinct i5 and i7 indices for dual‑indexed Illumina multiplexing.
Size Selection: Bead‑based purification returns ~300–600 bp libraries—optimal for Illumina® platforms.
Performance from peer‑reviewed literature: ClickSeq avoids fragmentation and enzymatic ligation, which dramatically reduces artifactual chimera formation and supports unbiased coverage across targets, with quality metrics on par with standard RNA/DNA‑seq approaches [1] [2].
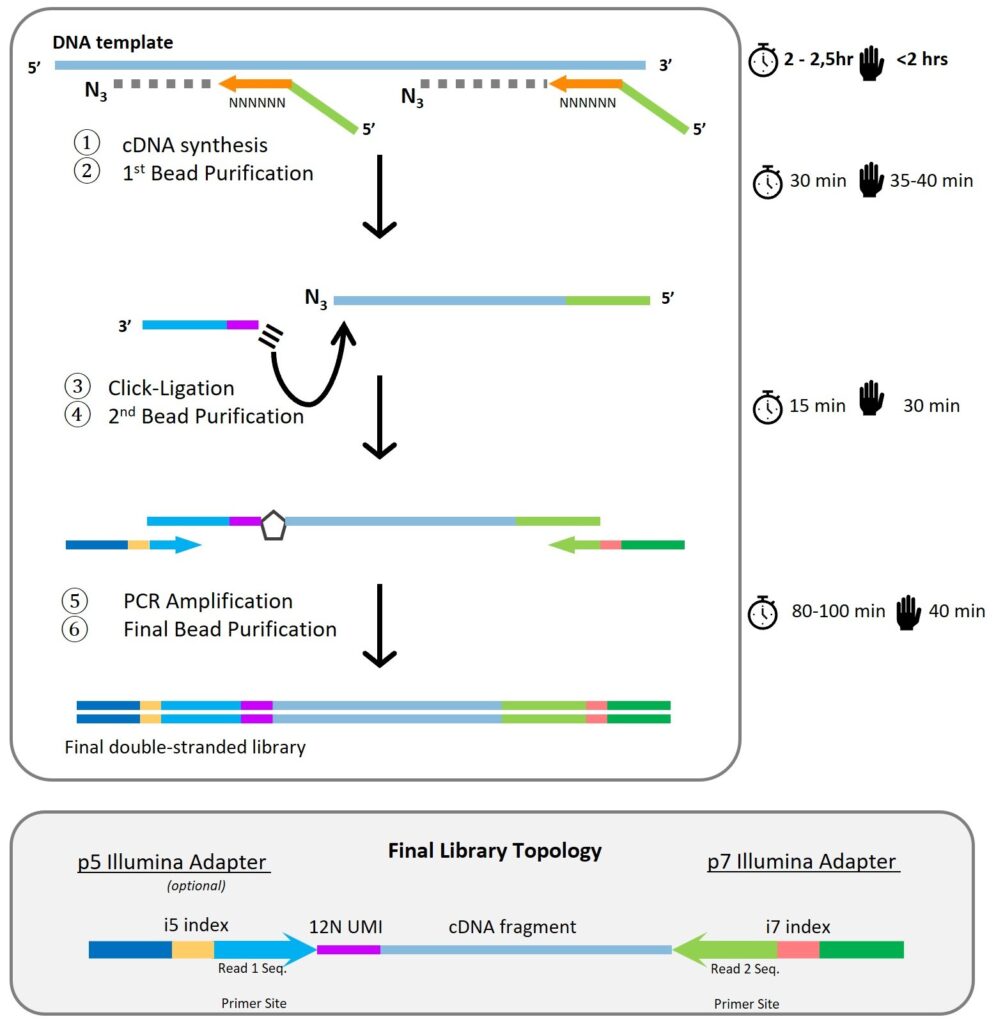
Figure 1: Flowchart of the ClickSeq method for library preparation.
Workflow Summary
- cDNA Synthesis: DNA is randomly primed and copied in the presence of azido-nucleotides, generating 3′-azido-terminated cDNA fragments.
- First Bead Purification: SPRI beads are used to purify cDNA fragments.
- Click-Ligation: Alkyne-functionalized adapters are covalently attached to the cDNA via click chemistry.
- Second Bead Purification: Removes reaction components, leaving adapter-flanked cDNA.
- PCR Amplification with Dual Indexing: Unique i5 and i7 index primers are used to amplify and barcode each sample.
- Final Bead Purification & Size Selection: Yields sequencing-ready libraries (optimal size: 300–600 bp).
Key Features & Advantages
- Dual Indexing: Supports both i5 and i7 indices for robust sample multiplexing and error correction.
- No DNA Fragmentation: Preserves genomic structure and reduces workflow complexity.
- Ultra-Low Chimera Rate: The stochastic termination and click-ligation process minimizes artifactual chimeras (as low as 3 per million reads).
- Low Bias & High Accuracy: Avoids enzymatic ligation and fragmentation-induced bias, improving variant detection and sequencing accuracy.
- Flexible Input Range: Works with 10 ng to 4 µg of DNA (optimal: >100 ng).
- Fast Protocol: Complete workflow in approximately 6 hours.
- Ready for Illumina Platforms: Generates libraries compatible with all major Illumina sequencers (NextSeq, NovaSeq, MiSeq, HiSeq).
- 12 Unique Dual-Index Primer Pairs: Enables multiplexing of up to 12 samples per kit.
Application Areas
- Human Population Genomics: Study genetic diversity and structural variation with high accuracy.
- Plant Breeding & Trait Discovery: Identify genetic traits and variations for crop improvement.
- Microbial Comparative Genomics: Analyze microbial diversity, pathogenicity, and antibiotic resistance.
Kit Components
- Click Primer Mix (CPM)
- Click Mix p5 (CM)
- Click Accelerant (CA)
- Click Catalyst (CC)
- Elution Buffers (EB1, EB2)
- Note: Enzymes (SuperScript III, RNase H, OneTaq Master Mix), SPRI beads are user-supplied. Dual Indexing Primers (i5 Index Primers (D501–D512), i7 Index Primers (D701–D712)) can be purchased here.
UMI: 12‑nt UMI incorporated in the read structure (per User Guide v1.0).
Library Topology
Final libraries include:
- Illumina p5 and p7 adapters
- Dual indices (i5 and i7)
- 4 nt UMI (unique molecular identifier)
- cDNA fragment
Technical Specifications
| Parameter | Specification |
| Input DNA | 10 ng – 4 µg; ≥100 ng recommended for complexity |
| Total time | ~6 hours including incubations |
| Library size | ~300–600 bp post‑cleanup |
| Indexing | Dual indexing (i5 + i7, 8‑nt) |
| UMI | 12 nt UMI captured at start of Read 1 (kit topology) |
| PCR cycles | Typical 12–21 (optimize by input) |
| Platforms | Illumina® MiSeq, NextSeq, NovaSeq, HiSeq |
| Throughput | 12 libraries per kit |
| ROU | Not for diagnostic procedures |
Data & Evidence (from the literature)
- Chimera suppression: “Fewer than 3 aberrant events per million reads” when avoiding fragmentation and ligation, enabling rare recombination analysis [1].
- Coverage & error: ClickSeq yields unbiased libraries with low error rates comparable to standard methods, confirming that CuAAC‑linked triazoles are compatible with downstream PCR and sequencing [1].
- Chemical rationale: The CuAAC reaction forms bio‑orthogonal triazole linkages that are read‑through by polymerases, supporting efficient library amplification and sequencing [2].
References
For detailed protocols and troubleshooting, refer to the https://www.clickseqtechnologies.com or contact ClickSeq Technologies at info@clickseq.com
LITERATURE
- Routh, A. et al. ClickSeq: Fragmentation‑Free Next‑Generation Sequencing via Click Ligation of Adaptors to Stochastically Terminated 3′‑Azido cDNAs. J. Mol. Biol. 427(16), 2610–2616 (2015). doi:10.1016/j.jmb.2015.06.011. (Free full text; reports <3 chimeras per 10⁶ reads and unbiased libraries with error rates comparable to standard methods.)
Link: https://europepmc.org/article/pmc/4523409
- Jaworski, E. & Routh, A. ClickSeq: Replacing Fragmentation and Enzymatic Ligation with Click‑Chemistry to Prevent Sequence Chimeras. In Next Generation Sequencing: Methods and Protocols, pp. 71–85 (2017/2018). doi:10.1007/978-1-4939-7514-3_6. (Protocol chapter detailing CuAAC/triazole biocompatibility and mechanism that underpins chimera suppression.)
Link: https://link.springer.com/protocol/10.1007/978-1-4939-7514-3_6
FAQ
-
What is DNA-ClickSeq Library Prep with Dual Indexing?
DNA-ClickSeq Library Prep with Dual Indexing is an advanced NGS library preparation kit that uses click chemistry for adapter ligation and supports dual-indexed sample barcoding. This kit is designed for researchers who want to buy a fast, reliable, and low-bias DNA library prep solution for Illumina sequencing platforms.
-
How can I purchase the DNA-ClickSeq Library Prep Kit?
You can order DNA-ClickSeq Library Prep with Dual Indexing directly from baseclick or through authorized distributors. Visit our online shop to buy DNA-ClickSeq kits or request a quote for bulk orders.
-
What are the main benefits of using this kit?
Fast workflow: Complete your NGS library prep in about 6 hours.
No DNA fragmentation required: Preserves sample integrity.
Dual indexing: Enables high-throughput multiplexing and accurate sample tracking.
Ultra-low chimera rate: Ensures high-quality sequencing data.
Compatible with Illumina platforms: Ready-to-sequence libraries for MiSeq, NextSeq, NovaSeq, and HiSeq.
-
What is included in the DNA-ClickSeq Library Prep with Dual Indexing kit?
Each kit contains all necessary reagents for 12 reactions, including click chemistry components. Enzymes and SPRI beads are user-supplied. For a full list, download the product datasheet or request a sample kit.
-
How do I get technical support or a user manual?
You can download the DNA-ClickSeq user manual from our website or contact our technical support team for troubleshooting and protocol optimization. Request technical support via our online form or email.
-
What DNA input amounts are supported?
The kit works with 10 ng to 4 µg of DNA. For best results, use at least 100 ng of high-quality DNA.
-
Can I use my own index primers?
Yes, as long as they are compatible with Illumina sequencing and match the adapter sequences used in the kit. For custom indexing solutions, contact our sales team.
-
How do I place a bulk order or request a quote?
To request a quote for bulk orders or to discuss custom solutions, please use our online quote request form or contact our sales department directly.
-
Is the kit available for international shipping?
Yes, we ship DNA-ClickSeq kits worldwide. Shipping options and costs are calculated at checkout. For distributor information, find a local distributor on our website.
-
What payment methods are accepted?
We accept major credit cards, PayPal, and institutional purchase orders. For more details, see our payment options during checkout.
-
Can I get a discount for academic or volume purchases?
Yes, special pricing is available for academic institutions and bulk purchases. Please contact us for a custom quote.
-
Is the DNA-ClickSeq Library Prep Kit for research use only?
Yes, this product is for research use only and is not intended for diagnostic or therapeutic applications.

