
Poly(A)-ClickSeq Library Prep Kit with dual indexing
Fragmentation-Free, Ligation-Free 3′ End RNA-Seq for Illumina® Platforms

| Size | Catalog No. | Price |
|---|---|---|
| 12 rxn | BCK-PACseqDual | € 396,00 |
Chemical Properties
-
Shelf Life
12 months unopened after receipt
-
Storage Conditions
– 20 °C
-
Physical State
kit system made of different components
-
CAS Number
n.a.
-
Preparation/Handling
please see user manual of the kit
Product Information
Technical Overview
Poly(A)-ClickSeq Library Prep with Dual Indexing is a next-generation sequencing (NGS) kit designed for precise 3′ end mapping of polyadenylated RNA, such as eukaryotic mRNAs. This innovative workflow leverages click chemistry for adapter ligation, eliminating the need for RNA fragmentation, selection, or enzymatic ligation. The kit enables robust dual-indexed sample tracking for high-throughput gene expression and alternative polyadenylation analysis.
How It Works:
- Azido-Termination: Total cellular RNA is reverse transcribed using an unanchored oligo-dT primer in the presence of azido-modified nucleotides (AzVTPs: AzATP, AzGTP, AzCTP). This stochastically terminates cDNA synthesis upstream of the poly(A) tail, generating cDNA fragments terminated with 3′-azido groups.
- Click-Ligation: Azido-terminated cDNA fragments are covalently linked to alkyne-functionalized sequencing adapters via copper-catalyzed azide–alkyne cycloaddition (CuAAC), also known as click chemistry. This chemical ligation is highly efficient and enzyme-free.
- Dual Indexing PCR Amplification: Click-ligated fragments are PCR-amplified using unique pairs of i5 and i7 index primers, enabling dual-indexed sample identification and multiplexing. The final libraries are fully compatible with Illumina sequencing workflows.
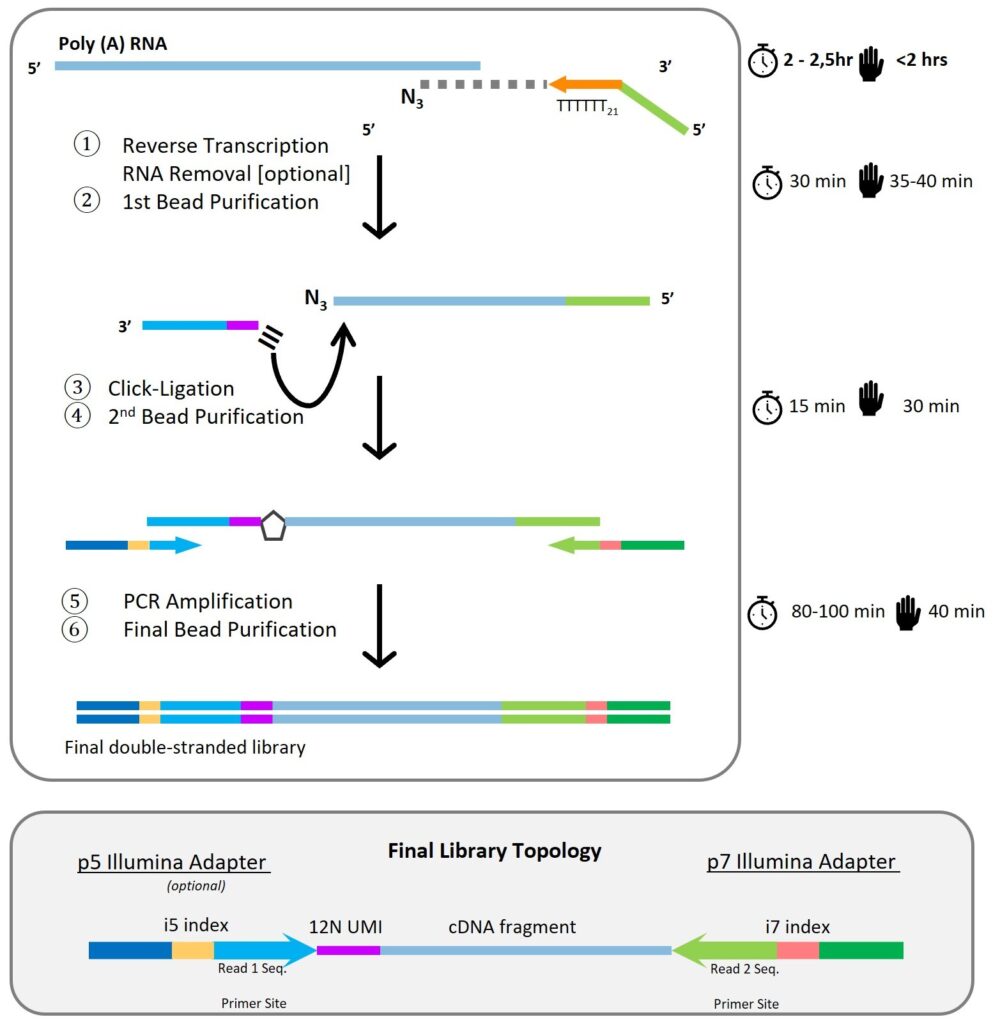
Workflow Summary
- Reverse Transcription: Total RNA is primed from poly(A) tails and copied in the presence of AzVTPs, generating 3′-azido-terminated cDNA fragments.
- Optional RNA Removal: RNase H can be used to remove template RNA.
- First Bead Purification: SPRI beads purify cDNA fragments.
- Click-Ligation: Alkyne-functionalized adapters are covalently attached to cDNA via click chemistry.
- Second Bead Purification: Removes reaction components, leaving adapter-flanked cDNA.
- PCR Amplification with Dual Indexing: Unique i5 and i7 index primers amplify and barcode each sample.
- Final Bead Purification & Size Selection: Yields sequencing-ready libraries (optimal size: 200–400 bp).
Key Features & Advantages
- 3′ End Targeting: Focuses on polyadenylated RNA for cost-efficient gene expression and poly(A) site mapping.
- No RNA Fragmentation or Selection: Directly uses crude total RNA, simplifying sample prep and reducing bias.
- Ultra-Low Chimera Rate: Click chemistry and stochastic termination minimize artifactual chimeras.
- Dual Indexing: Supports both i5 and i7 indices for robust sample multiplexing and error correction.
- Flexible Input Range: Works with 30 ng to 4 µg total RNA (optimal: 1 µg).
- Fast Protocol: Complete workflow in approximately 6 hours.
- Ready for Illumina Platforms: Generates libraries compatible with all major Illumina sequencers (NextSeq, NovaSeq, MiSeq, HiSeq).
- 12 Unique Dual-Index Primer Pairs: Enables multiplexing of up to 12 samples per kit.
Application Areas
- Gene Expression Profiling: Cost-effective quantification of transcript abundance.
- Alternative Polyadenylation Studies: High-resolution mapping of poly(A) sites.
- Differential Poly(A) Site Usage: Supports studies of mRNA processing and regulation.
- High-Throughput Screening: Dual indexing enables efficient multiplexing for large cohorts.
Kit Components
- PAC Primer Mix (PPM)
- Click Mix p5 (CM)
- Click Accelerant (CA)
- Click Catalyst (CC)
- Elution Buffers (EB1, EB2)
- Note: Enzymes (SuperScript III, RNase H, OneTaq Master Mix), SPRI beads are user-supplied. Dual Indexing Primers (i5 Index Primers (D501–D512), i7 Index Primers (D701–D712)) can be purchased here.
Library Topology
Final libraries include:
- Illumina p5 and p7 adapters
- Dual indices (i5 and i7)
- 4 nt semi-UMI (unique molecular identifier)
- cDNA fragment
- Poly(A) stretch
Technical Data
- High Sensitivity and Specificity: Poly(A)-ClickSeq accurately maps polyadenylation sites with single-nucleotide resolution, enabling robust detection of alternative polyadenylation events.
- No Fragmentation Required: The protocol avoids RNA fragmentation, reducing bias and preserving transcript integrity.
- Low Chimera Formation: The click chemistry approach results in extremely low levels of chimeric reads, supporting high-confidence data analysis.
- Direct Use of Total RNA: No need for poly(A) selection or ribosomal RNA depletion; the kit works directly from total cellular RNA.
- Efficient Library Construction: Libraries are generated with high complexity and uniform coverage, suitable for differential gene expression and poly(A) site usage studies.
Optimized for Alternative Polyadenylation Analysis: Enables quantitative profiling of poly(A) site usage across samples and conditions.
Compatibility & Guidelines
Compatible with all Illumina sequencing platforms.
Not recommended for multiplexing with libraries prepared by other methods due to differences in library topology and poly(A) tract diversity.
For research use only. Not intended for diagnostic or therapeutic applications.
Technical Specifications
| Parameter | Specification |
| Input RNA | 30 ng – 4 µg (optimal: >100ng) |
| Total time | ~6 hours including incubations |
| Library size | ~200–400 bp post-cleanup |
| Indexing | Dual indexing (i5 + i7, 8‑nt) |
| UMI | 4 nt semi-UMI |
| PCR cycles | Typical 12–21 (optimize by input) |
| Platforms | Illumina® MiSeq, NextSeq, NovaSeq, HiSeq |
| Throughput | 12 libraries per kit |
| ROU | Not for diagnostic procedures |
References
For detailed protocols and troubleshooting, refer to the user manual or visit ClickSeq Technologies.
LITERATURE
Routh, A., et al. (2017). Poly(A)-ClickSeq: click-chemistry for next-generation 3′-end sequencing without RNA enrichment or fragmentation. Nucleic Acids Research, 45(12), e112. https://doi.org/10.1093/nar/gkx426
FAQ
-
What is Poly(A)-ClickSeq Library Prep with Dual Indexing?
Poly(A)-ClickSeq is a next-generation sequencing (NGS) kit for preparing Illumina-compatible libraries that target the 3′ ends of polyadenylated RNA (e.g., mRNA). It uses click chemistry for adapter ligation and supports dual-indexed sample barcoding for high-throughput multiplexing.
-
What are the main advantages over conventional RNA-seq kits?
No RNA fragmentation or selection required—works directly from total RNA.
Ultra-low chimera rate due to click chemistry and stochastic termination.
High sensitivity and specificity for poly(A) site mapping.
Fast, streamlined workflow (~6 hours).
Robust dual indexing (i5/i7) for accurate sample tracking.
-
What RNA input amounts are compatible?
The kit works with 30 ng to 4 µg total RNA. For best results, use 1 µg of high-quality RNA.
-
Which sequencing platforms are compatible?
Libraries are fully compatible with all Illumina sequencing platforms (MiSeq, NextSeq, NovaSeq, HiSeq).
-
Can I use my own index primers?
Yes, as long as they are compatible with Illumina sequencing and match the adapter sequences used in the kit.
-
Do I need to enrich for poly(A) RNA or deplete rRNA?
No. The kit is designed to work directly from crude total RNA without poly(A) selection or rRNA depletion.
-
Is RNA fragmentation required?
No. Poly(A)-ClickSeq does not require RNA fragmentation, reducing bias and preserving transcript integrity.
-
How long does the workflow take?
The complete protocol can be finished in approximately 6 hours, including hands-on and incubation times.
-
What is the typical library size?
The protocol yields libraries in the 200–400 bp range, optimal for Illumina sequencing.
-
Can I multiplex Poly(A)-ClickSeq™ libraries with libraries from other kits?
It is not recommended, as differences in library topology and poly(A) tract diversity may affect sequencing quality and data output.
-
What is the UMI and how is it used?
Each library molecule contains a 4 nt semi-unique molecular identifier (semi-UMI) for accurate deduplication and error correction during data analysis.
-
What quality control steps are recommended?
Quantify libraries using Qubit or BioAnalyzer, and check size distribution before pooling and sequencing.
-
Is the kit for research use only?
Yes, Poly(A)-ClickSeq Library Prep Kit is for research use only and not intended for diagnostic or therapeutic applications.
-
Where can I find the detailed protocol and troubleshooting guide?
Refer to the user manual included with the kit or visit the ClickSeq Technologies website for the latest documentation and support.

